Veröffentlicht 8. Februar 2018 von Melissae Fellet
Der 3D-Code des Genoms
Der genetische Code besteht aus einer Reihe von Buchstaben, welche die Anleitungen für das normale Wachstum, die Reparatur und die routinemäßigen Abläufe in Zellen buchstabieren. Inzwischen deutet vieles darauf hin, dass die verknäuelte DNA-Struktur einen zweiten Code enthält. Möglicherweise steuern Position und Packungsdichte von Nukleinsäuren, welche Instruktionen zu einem bestimmten Zeitpunkt zugänglich und aktiv sind. Eine beeinträchtigte Genomstruktur könnte unter anderem für Krankheiten wie Krebs und Missbildungen verantwortlich sein.
Damit sie in das Innere einer Zelle passt, vollbringt die DNA eine unglaubliche akrobatische Meisterleistung: Zwei Meter Material werden in einen nur ein paar Mikrometer messenden Zellkern gepresst. Die DNA verdichtet sich eigenständig, indem sie sich zunächst um Histonproteine wickelt und so eine Nukleosomkette bildet, die wie eine Perlenschur aussieht. Anschließend wickeln sich die Nukleosomen zu Chromatinfasern auf, die sich wie Spaghetti in einem Topf verknäueln.
Zur Aufdeckung des strukturellen genetischen Codes untersuchen Forscher das Chromatin von der Nukleotidsequenz bis zur Organisation eines gesamten Genoms. Aufgrund der Entwicklung mikroskopischer Verfahren zur verbesserten visuellen Darstellung der detaillierten Chromatinstruktur – sogar in lebenden Zellen – ist man heute noch besser in der Lage, den Zusammenhang zwischen Strukturveränderungen und Genexpression bzw. Zellfunktion zu erforschen. Die heute möglichen Aufnahmen der Chromatinstruktur helfen bei der Beantwortung einiger der größten Fragen der Genombiologie.
Chromatinkompartimente
Eine vorherrschende Theorie bezüglich der Chromatinstruktur besagt, dass sich Nukleosomen zu 30 nm dicken Fasern aufwickeln, die sich dann zu zunehmend größeren Strukturen zusammenlagern und schließlich Chromosomen bilden. Der Beleg hierfür sind Beobachtungen, nach denen in gereinigter Form aus Zellen gewonnene DNA und Nukleosomen 30 nm bzw. 120 nm dicke Fasern bilden.
In der Arbeitsgruppe von Clodagh O’Shea am Salk Institute for Biological Studies fragte man sich, wie Chromatin wohl in intakten Zellen aussieht. 2017 entwickelten die Forscher ein Verfahren zur Sichtbarmachung von Chromatin in intakten menschlichen Zellen, die ruhten bzw. sich teilten. Die Forscher beschichteten die Zell-DNA mit einem Material, das Osmiumionen absorbiert, was dazu führt, dass die Nukleinsäure einen Elektronenstrahl besser streuen kann und dadurch in einer elektronenmikroskopischen Aufnahme sichtbar wird. Als Nächstes wendeten sie ein modernes elektronenmikroskopisches Verfahren an, bei dem die Proben schräg in einen Elektronenstrahl gehalten werden und das dreidimensionale Strukturinformationen liefert. Die Forscher stellten fest, dass Chromatin in Form einer semiflexiblen, 5 bis 24 nm dicken Kette vorlag, die in einigen Abschnitten der Zelle dicht und in anderen locker gepackt war.
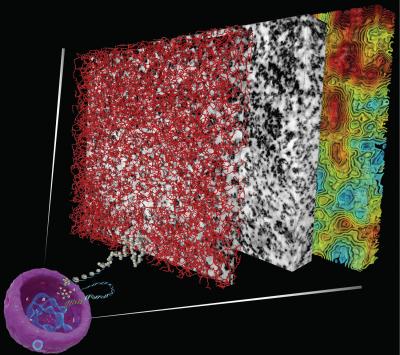
“Wir zeigen, dass Chromatin keine klaren Strukturen höherer Ordnung bilden muss, damit es in den Zellkern passt”, so O’Shea. „Es ist die Packungsdichte, die möglicherweise die Zugänglichkeit des Chromatins verändert und begrenzt, indem sie eine lokale und globale Strukturbasis für die Integration verschiedener Kombinationen von DNA-Sequenzen, Nukleosomvariationen und -modifikationen in den Nukleus zur minutiösen Feinabstimmung der funktionellen Aktivität und Zugänglichkeit unserer Genome bereitstellt.“
Neben der Packungsdichte stellt die Position eine weitere Komponente der strukturellen Organisation des Chromatins dar. Es ist bereits seit 30 Jahren bekannt, dass Chromatin Schleifen bildet und so Gene näher zu Sequenzen hinzieht, die ihre Expression regulieren. Der Biologe Job Dekker und seine Mitarbeiter von der University of Massachusetts Medical School in Worcester haben verschiedene molekularbiologische Techniken zur Identifizierung benachbarter, 200.000 bis eine Million Basen langer Chromatinabschnitte entwickelt. Bei einer dieser Techniken, dem sogenannten Hi-C, wird die Chromatinstruktur anhand ihrer Sequenz kartiert.
Bei dem Hi-C-Verfahren wird die Nukleinsäure zunächst mit nahe beieinander liegenden Chromatinabschnitten chemisch vernetzt. Anschließend wird das vernetzte Chromatin mit Hilfe von Enzymen zerschnitten und die losen Enden werden mit einem modifizierten Nukleotid markiert. Danach werden nur vernetzte Fragmente wieder miteinander verbunden. Schließlich isolieren die Forscher die Chromatinfragmente, sequenzieren sie und ordnen die Sequenzen ihrer Position im Gesamtgenom einer Zelle zu.
Im Jahr 2012 identifizierten Bing Ren und seine Mitarbeiter von der University of California, San Diego School of Medicine mittels Hi-C Chromatinregionen, die sie als topologisch assoziierte Domänen (TAD) bezeichneten. Gene in ein- und derselben TAD treten stärker miteinander in Wechselwirkung als mit Genen in anderen TAD, und Domänen, in denen eine aktive Transkription stattfindet, besetzen andere Positionen in einem Zellkern als ruhende Domänen. Veränderte Sequenzen innerhalb einer TAD können zu Krebs und Gliedmaßenfehlbildungen bei Mäusen führen.
Man geht davon aus, dass durch einen Proteinanker gezogene Chromatinschleifen die Grundeinheit einer TAD bilden. In modernen Computermodellen der Chromatinfaltung lassen sich mittels Hi-C beobachtete Wechselwirkungen des Chromatins bei der Schleifenbildung simulieren. Genomforscher sind sich jedoch nach wie vor nicht sicher, welche Proteine an der Schleifenbildung beteiligt sind. Die Beantwortung dieser Frage betrifft eine grundlegende Eigenschaft der DNA-Faltung und könnte auf einen zellulären Krankheitsmechanismus infolge von Mutationen in einem Schleifenverankerungsprotein deuten.
Superauflösende Mikroskopie
Moderne Methoden der optischen Mikroskopie, die auf einem Verfahren basieren, für das der Nobelpreis für Chemie 2014 verliehen wurde, liefern ebenfalls Informationen darüber, wie bis zu 100 Basen lange Chromatinregionen die Zellfunktion beeinflussen könnten. Die superauflösende Fluoreszenzmikroskopie verstärkt die Auflösung von Lichtmikroskopen, so dass die Beugungsgrenze bei mehr als 300 nm liegt. Bei dieser Technik werden fluoreszierende Moleküle mit Hilfe eines Lichtimpulses angeregt. Anschließend wird das Licht, das von nicht zentral im Anregungsstrahlengang befindlichen Molekülen emittiert wird, mittels verschiedener Kunstgriffe unterdrückt. Auf diese Weise lassen sich Aufnahmen eines einzelnen fluoreszierenden Moleküls erzeugen.
Biologische Moleküle können jedoch zahlreiche fluoreszierende Marker aufweisen, was die Lokalisation eines Einzelmoleküls erschwert. Mit Hilfe von fluoreszierenden Markern, die sich an- und ausschalten lassen, aktivieren und deaktivieren Forscher Moleküle in bestimmten Regionen zu bestimmen Zeitpunkten. Dann verbinden sie die Aufnahmen und erfassen die Positionen aller Fluoreszenzmarkierungen.
Xiaowei Zhuang und seine Mitarbeiter von der Harvard University verfolgten mittels superauflösender Mikroskopie, wie sich die Chromatinpackung basierend auf epigenetischen Modifikationen veränderte. Ihre Methode lieferte Aufnahmen im Kilo- bis Megabasen-Maßstab; diese Auflösung liegt zwischen der reiner Sequenzinformationen und der weitreichender Wechselwirkungen, die durch Hi-C darstellbar sind, wie zum Beispiel Informationen in Bezug auf Genregulation und -transkription. Dieses Verfahren eröffnet auch die Möglichkeit einer bildgebenden Darstellung von Strukturen im Nanometerbereich in lebenden Zellen.
Strukturwörterbuch
Derzeit arbeiten Forscher auf der ganzen Welt an einem Wörterbuch des strukturellen genetischen Codes in Raum und Zeit und wenden dabei eine Vielzahl von Methoden zur Erfassung statischer und dynamischer Zellveränderungen an. In dem von den National Institutes of Health gegründeten 4D Nucleome Network und dem vom Europäischen Forschungsrat ins Leben gerufenen 4D Genome Project wird das Vokabular der DNA-Strukturelemente ermittelt und untersucht, welche Auswirkungen diese Struktur auf die Genexpression hat. Man ist zudem neugierig, wie sich die Chromatinstruktur im Verlauf der normalen Entwicklung sowie bei Krankheiten wie Krebs und vorzeitigem Altern verändert. Auch wenn bereits viele grundlegende Fragen geklärt werden konnten, so bleibt trotz allem noch viel zu entdecken.