
Ingredients for a Career in Science
Kamaljit Moirangthem has been living and researching in seven countries so far. He has just started his new position in Bergen, Norway. So, he is quite experienced to talk about mobility and mentoring, and about sustainable food production.

World Tuberculosis Day: A Closer Look at the Challenges in Eradicating TB
24 March is the annual World Tuberculosis Day, announced by the WHO. TB is the world’s leading cause of death from infectious disease.
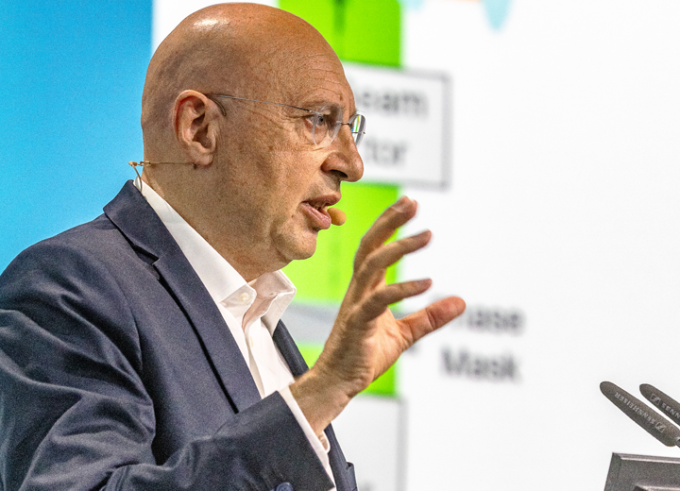

Breaking the Diffraction Limit
During an interview in Lindau in occasion of #LINO23, Stefan Hell talked about his past work, how the landscape has changed for young researchers since he started out in science, and why he attends the Lindau Nobel Laureate Meetings almost every year.
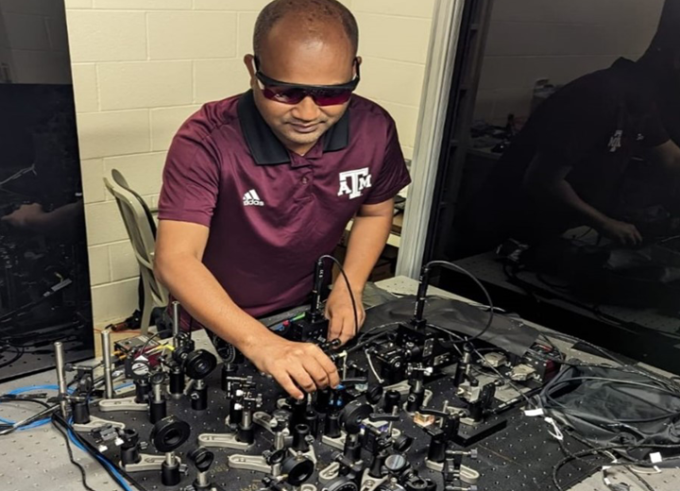

Physics Will Love You!
#LINO19 Alumnus Tanvir Islam Rajib is working in the field of quantum physics at Texas A&M University. Quantum physics and quantum technologies are, among others, key topics of #LINO24.

Women in Research #LINO23: Maria Bartosova
We are closing the #LINO23 "Women in Research" series with Maria Bartosova whose research helps children suffering from chronic kidney diseases.

Women in Research #LINO23: Cornelia Schwayer
Cornelia Schwayer studies the intestinal organoid, a self-organizing system that forms a complex tissue from a single cell, to understand how organs regenerate.
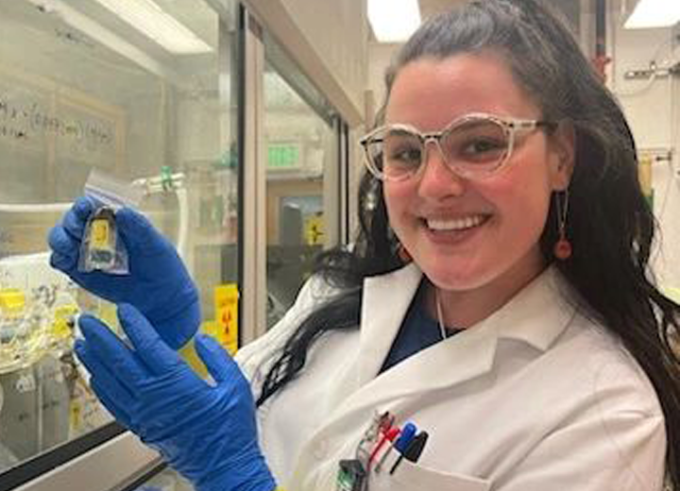

Women in Research #LINO23: Alexia Cosby
As a chemist, Alexia Cosby is inspired by the ability to build intricate, complicated compounds from scratch which have incredible powers to image, treat, and irradicate various cancers and diseases.
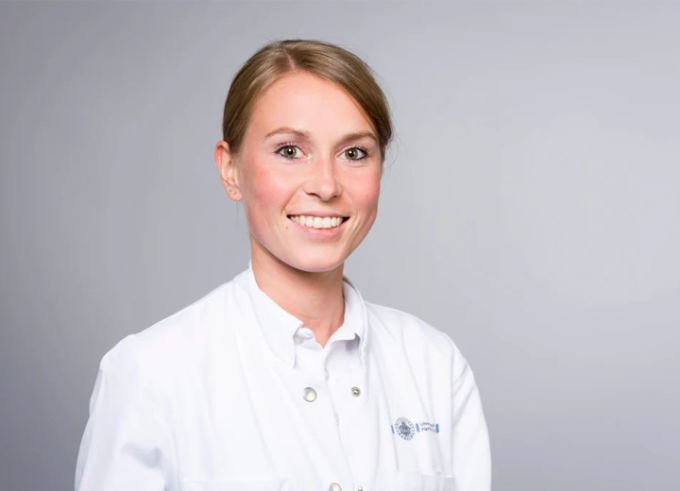

Women in Research #LINO23: Märit Jensen
Märit Jensen, Lindau Alumna 2023, focuses on imaging-based clinical stroke research and the study of brain reorganisation after stroke and cerebrovascular diseases.
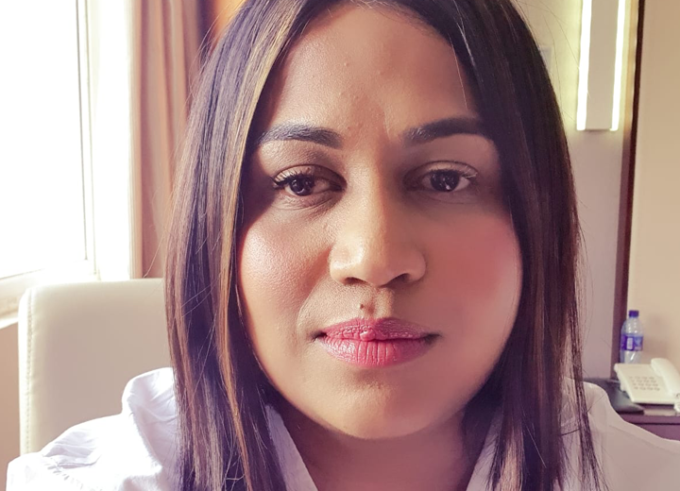

Make a Difference in the World!
For Zaynah Dhunny, the participation in the 66th Lindau Nobel Laureate Meetingwas a milestone that changed her career direction.

Oppenheimer: A Cinematic Journey Through the Lives of Legendary Physicists
One of the most successful movies in 2023 was the biopic "Oppenheimer". A must-seen for everyone interested in Physics. Andrei Mihai presents the leading physicists who are featured in the film.

Women in Research #LINO23: Alexandrina Vasilichi
Alexandrina Vasilichi investigates the computational mechanisms by which interoceptive signals control affective expectations and affective learning processes.
Der Nobelpreis für Physiologie/Medizin 2023: Ein neuer Blick auf Impfstoffe
Der Nobelpreis für Physiologie oder Medizin 2023 geht an Katalin Karikó und Drew Weissman, die die Grundlage für die Erfindung von mRNA-Impfstoffen gelegt haben. Ihre Forschung ermöglichte es, COVID-19-Impfstoffe schnell zu entwickeln.











